Identify clusters of cells from a shared nearest-neighbor graph. More...
#include "../utils/macros.hpp"#include "BuildSnnGraph.hpp"#include <vector>#include <algorithm>#include "igraph.h"#include "igraph_utils.hpp"
Include dependency graph for ClusterSnnGraph.hpp:
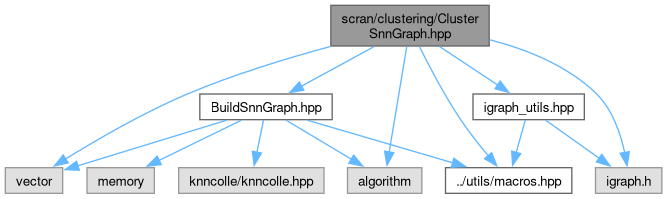
Go to the source code of this file.
Classes | |
| class | scran::ClusterSnnGraphMultiLevel |
| Multi-level clustering on a shared nearest-neighbor graph. More... | |
| struct | scran::ClusterSnnGraphMultiLevel::Defaults |
| Default parameter settings. More... | |
| struct | scran::ClusterSnnGraphMultiLevel::Results |
| Result of the igraph multi-level community detection algorithm. More... | |
| class | scran::ClusterSnnGraphWalktrap |
| Walktrap clustering on a shared nearest-neighbor graph. More... | |
| struct | scran::ClusterSnnGraphWalktrap::Defaults |
| Default parameter settings. More... | |
| struct | scran::ClusterSnnGraphWalktrap::Results |
| Result of the igraph Walktrap community detection algorithm. More... | |
| class | scran::ClusterSnnGraphLeiden |
| Leiden clustering on a shared nearest-neighbor graph. More... | |
| struct | scran::ClusterSnnGraphLeiden::Defaults |
| Default parameter settings. More... | |
| struct | scran::ClusterSnnGraphLeiden::Results |
| Result of the igraph leiden community detection algorithm. More... | |
Namespaces | |
| namespace | scran |
| Functions for single-cell RNA-seq analyses. | |
Detailed Description
Identify clusters of cells from a shared nearest-neighbor graph.