Model the per-gene variance from log-count data. More...
#include "../utils/macros.hpp"#include "tatami/tatami.hpp"#include "../utils/vector_to_pointers.hpp"#include "../utils/blocking.hpp"#include "../utils/average_vectors.hpp"#include "FitVarianceTrend.hpp"#include "blocked_variances.hpp"#include <algorithm>#include <vector>#include <limits>
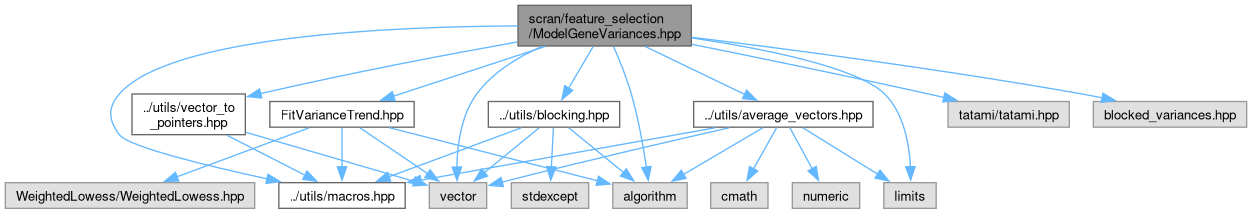
Include dependency graph for ModelGeneVariances.hpp:
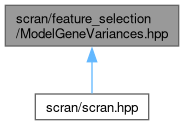
This graph shows which files directly or indirectly include this file:
Go to the source code of this file.
Classes | |
| class | scran::ModelGeneVariances |
| Compute and model the per-gene variances in log-expression data. More... | |
| struct | scran::ModelGeneVariances::Defaults |
| Default parameters for variance modelling. More... | |
| struct | scran::ModelGeneVariances::Results |
| Results of variance modelling without blocks. More... | |
| struct | scran::ModelGeneVariances::BlockResults |
| Results of variance modelling with blocks. More... | |
Namespaces | |
| namespace | scran |
| Functions for single-cell RNA-seq analyses. | |
Detailed Description
Model the per-gene variance from log-count data.