Set common macros used through libscran. More...
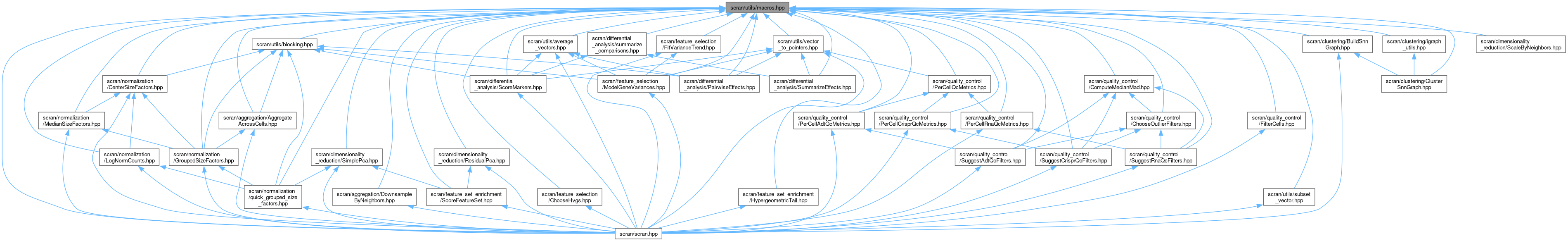
Go to the source code of this file.
Detailed Description
Set common macros used through libscran.
The SCRAN_CUSTOM_PARALLEL macro can be set to a function that specifies a custom parallelization scheme. This function should be a template that accept three arguments:
njobs, an integer specifying the number of jobs.fun, a lambda that accepts two arguments,startandend.nthreads, an integer specifying the number of threads to use.
The function should split [0, njobs) into any number of contiguous, non-overlapping intervals, and call fun on each interval, possibly in different threads. The details of the splitting and evaluation are left to the discretion of the developer defining the macro. The function should only return once all evaluations of fun are complete.
If SCRAN_CUSTOM_PARALLEL is set, the following macros are also set (if they are not already defined):
This ensures that any custom parallelization scheme is propagated to all of libscran's dependencies. If these libraries are used outside of libscran, some care is required to ensure that the macros are consistently defined through the client library/application; otherwise, developers may observe ODR compilation errors.